- Home
- About
- Contact
- Darkwood hideout 3 do you not regen health at night here
- Best mac screen grab recording
- Minecraft pocket edition free demo
- Iexplorer software coupons
- Battle of los angeles movie crossover
- Catz 5 ptzfiles
- Best free task manager 2019
- Betaflight flight controller failed to open serial port
- Different file formats for ucsc genome browser
- Apple drawing pad for macbook air
- How to use google pinyin installer
- Download pdf to music pro with crack
- Neat video 4-6 crack torrent download
- Node js windows set debug
- Wiz khalifa taylor allderdice mixtape tracklist
- Pyspark on windows 10
- Project 2016 for office 365
- Minecraft server list 1-5 2
- Acrok video converter ultimate crack
- Project table of contents example
- Heaven by bryan adams on guitar tab pdf
- Checkpoint vpn client mac download
- Microsoft office for educators for mac
- Epson wf 2540 printer error message
- Duplicate photo finder for mac
- Broadcom 802-11g network adapter driver windows 7
- Free online video editor and maker and free donwloasd
- Stop motion animation software chrome os
- How to enter customer number in final draft 10
- Titanium tv download update
- Toshiba printer drivers e studio 2050c
- Kanto karaoke software package
- How to clean up mac files
- Minecraft pe shaders how to install
- Macbook pro cable to 3 rca cable tv
- Do i want the new version of skype
- Tge apple mac 27
- Used apple laptop 2016
- Son of the mask free full movie
- Where to download microsoft office home and student 2007
- Ableton live 10 key mappings
- Best keynote presentation templates - training 2018
- #Different file formats for ucsc genome browser how to#
- #Different file formats for ucsc genome browser install#
- #Different file formats for ucsc genome browser code#
- #Different file formats for ucsc genome browser download#
The last step assumes that your ~/public_html/ directory is accessible from the internet. bedToBigBed bedExample.txt myBigBed.bb The steps areĮxplained in more detail in the following sections on this page: (swap macOSX for linux for an Apple environment). The following UNIX commands create one on a Linux machine Note that the bedToBigBed utility uses a substantial amount of memory:Īpproximately 25% more RAM than the uncompressed BED input file. See this wiki page for help in selecting the graphing track data format that is most
#Different file formats for ucsc genome browser how to#
See Example #3 below for an example of how to build an additional Hosting section of the Track Hub Help documentation.Īdditional indices can be created for the items in a bigBed file to support item search in track If you do not haveĪccess to a web-accessible server and need hosting space for your bigBed files, please see the Server (http, https, or ftp), not on the UCSC server, and only the portion that is needed for theĬurrently displayed chromosomal position is locally cached as a "sparse file". The bigBed file remains on your local web-accessible Regular BED when working with large data sets. Because of this, bigBed has considerably faster display performance than Those portions of the files needed to display a particular region are transferred to the Genomeīrowser server. The main advantage of the bigBed files is that only The resulting bigBedįiles are in an indexed binary format. BigBed files are createdįrom BED type files using the program bedToBigBed.
#Different file formats for ucsc genome browser code#
Once the code is finished, check *BooBED_All.txt for predicted TFBS branch-of-origin.The bigBed format stores annotation items that can be either a simple or a linked collection ofĮxons, much as BED files do. boo -s data/CTCF/hg19_multiz_Broad_ -r data/CTCF/chip_genCoord_Broad_ -t data/tree_file.nh -x hg19 -k 100 -w -100:100 -a GGGGCKC -q CTCF_kmer.txt boo -m f -s data/CTCF/hg19_multiz_Broad_ -r data/CTCF/chip_genCoord_Broad_ -t data/tree_file.nh -x hg19 -k 100 -w -100:100 -a GGGGCKC -q CTCF_kmer.txt Output files will be written into the same folder as multiple sequence alignment file. Then create a link pointing to those files or copy files to that folder. Once multiple sequence alignment and kmer list are generated, create a new folder (e.g. The meaning of parameter is same as main program BOO. boo_pwm -m w -s data/hg19/CTCF/hg19_multiz_Broad_ -r data/hg19/CTCF/chip_genCoord_Broad_ -w -100:100 -k 100 -t data/tree_file.nh -x hg19 -a GGGGCKC So please include both kmer and its reverse complement.) User can also provide a customer kmer list generated by any other methods (*BOO will not automatically generate reverse complement of kmer. Along with BOO program, we also provide another tools called BOO_pwm to generate PWM motif profile from ChIP-seq peak region and output a kmer list. MafsInRegion chip_genCoord_Broad_ hg19_multiz_Broad_ Genome_UCSC/Human/hg19/multiz46way/*.mafįor a known TF, in order to reduce the calculation time, we use a list of re-computed kmer list to search for TFBS (see example kmer file here). Use mafsInRegion to extract multiple sequence alignment within ChIP-seq peak. Genome_UCSC/Human/hg19/multiz46way/) and uncompress them. Other required files can be generated using the following scripts/tool.ĭownload human multiz46way alignments file from UCSC Genome Browser, put those files in folder (e.g.
#Different file formats for ucsc genome browser download#
Example ChIP-seq peak file can be download here. A threshold will be used to filter peaks with score below user specifiec cutoff(-k). , in which is TF name, is peak rank and is an interger score indicating the peak height.

A ChIP-seq peak file in four column bed format is required to run BOO.
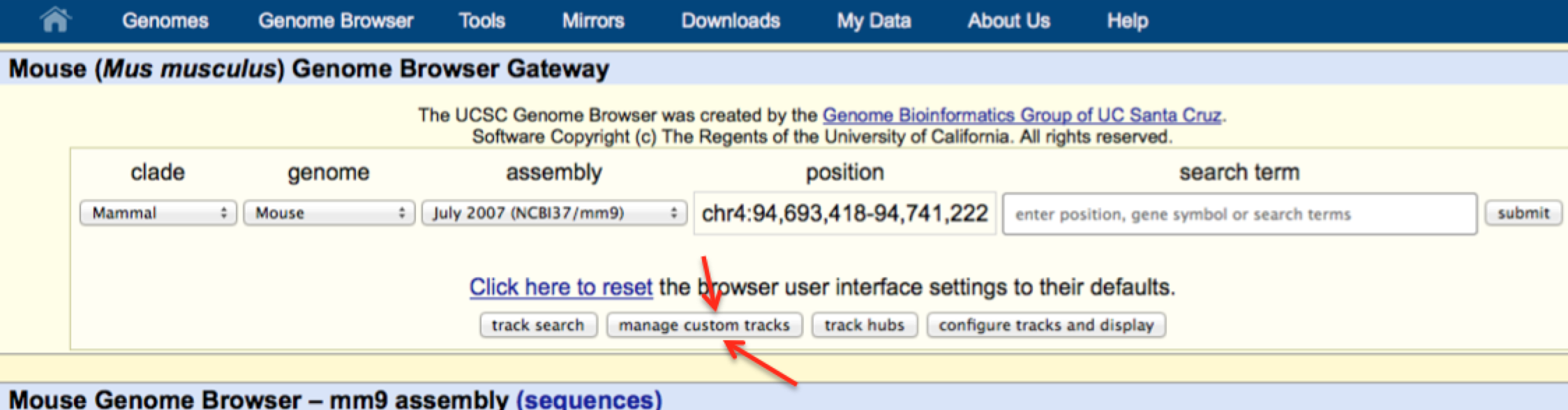
Executable binary programs were also provided.
#Different file formats for ucsc genome browser install#
The fourth column indicates the predicted branch-of-origin of TFBSĭownload the package, uncompressed the file using tar -zxvf boo_0.1.0.tar.gz and follow the instruction in README.txt to install the package. Predicted branch-of-orgin of TFBS in bed file format.
